In one of our recent papers, we used an AI model to predict treatment response and overall survival in intensively treated acute myeloid leukemia (AML). The article was well received by the community rendering it one of the four most downloaded papers in Haematologica for 2023, so if you haven’t checked it out already, now is the time!
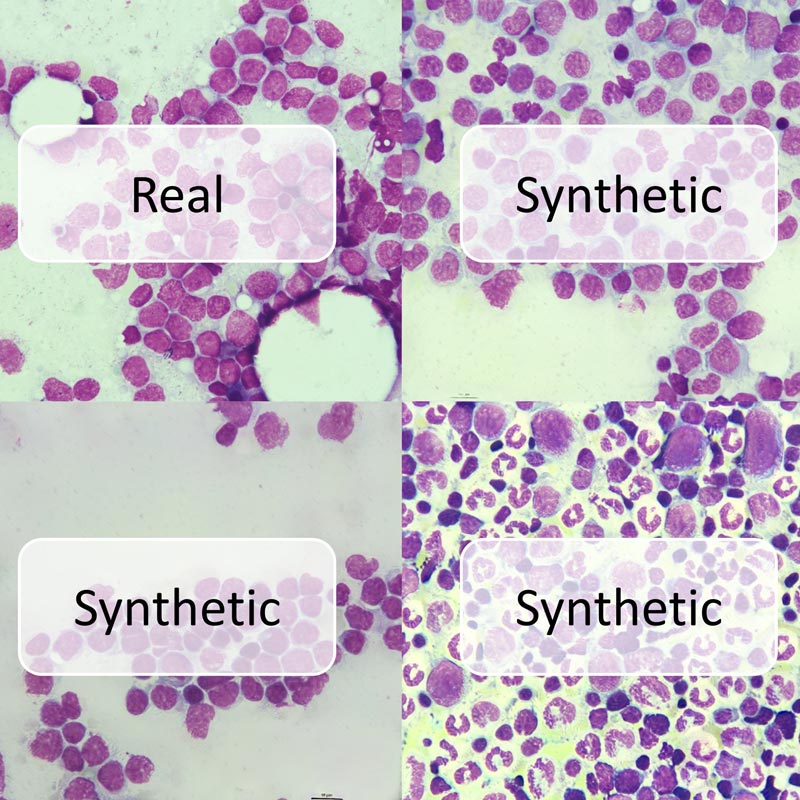
Back then, we used NGS panel data in conjunction with clinical information to train the classifier. Apart from the ‚usual suspects‘ in AML genetics, we were pretty surprised that the AI picked up on a gene that was kind-of coincidentally included in the panel, IKZF1, and identified it as a strong predictive marker for outcome. IKZF1 is a tumor supressor which is well known to negatively affect outcome in acute lymphoblastic leukemia, but it was not well studied in AML.
After the AI has led us on the track, we had a closer at IKZF1 in the lab. We were able to identify a mutational hotspot in AML that abrogates IKZF1’s ability to bind DNA. Further, IKZF1 turned out to be a strong predictive marker of decreased treatment response, and poor event-free, relapse-free, and overall survival in multivariable analyses.
This tale underlines the power of AI in genomics: AI can help us identify genes that matter for an in-depth analysis of disease biology.
Check out our new paper in Nature Leukemia about IKZF1 in AML:
Papers:
Eckardt JN, Stasik S, Röllig C, et al. Mutated IKZF1 is an independent marker of adverse risk in acute myeloid leukemia. Leukemia. 2023;37(12):2395-2403. doi:10.1038/s41375-023-02061-1
https://www.nature.com/articles/s41375-023-02061-1
Eckardt JN, Röllig C, Metzeler K, et al. Prediction of complete remission and survival in acute myeloid leukemia using supervised machine learning. Haematologica. 2023;108(3):690-704. Published 2023 Mar 1. doi:10.3324/haematol.2021.280027
https://www.haematologica.org/article/view/haematol.2021.280027